Innovation and Collaboration Grants
Salk’s Innovation and Collaboration Grants are a unique, philanthropically funded grant program launched through the vision and generosity of Joan and Irwin Jacobs. These internal awards empower our faculty to pursue high-risk, high-reward ideas that don’t quite fit the mold of traditional federal grants.
The Innovation Grant Program was launched in 2006 to fund early-stage, out-of-the-box ideas that hold significant promise. Awarded semi-annually by peer review, Innovation Grants are critical for catalyzing emerging science with the power to redefine the future.
The Collaboration Grant Program was launched in 2019 to foster new partnerships between Salk scientists. By encouraging collaboration across multiple labs, the Institute stands on its belief that the biggest scientific breakthroughs emerge from the cross-pollination of ideas between disparate fields of research.
What the Jacobses understood is that all breakthroughs start somewhere. Now also supported by donors Sarah and Jay Flatley, Richard Heyman and Anne Daigle, and the NOMIS Foundation, these early-stage grants have the potential to birth new fields of science and inspire life-changing advances.
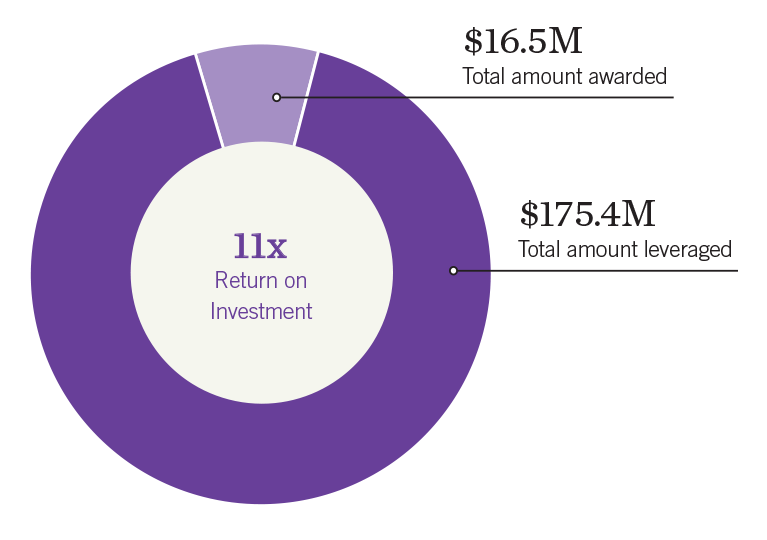
Since 2006, more than $16 million in seed funding has enabled Salk researchers to secure more than $175 million in follow-up federal, foundation, and industry grants—a remarkable 11-fold return on investment.
2025 Innovation and Collaboration Grants
Innovation Grant
When we chew, speak, or drink, the tongue moves with exquisite speed and precision inside the mouth, working harmoniously with our breathing and ongoing jaw movements. Disrupted tongue control is common after brain injury (e.g., stroke) and neurodegenerative disease (e.g., amyotrophic lateral sclerosis, Parkinson’s disease), often leading to severe disability or even death. Despite its vital importance, we lack a comprehensive understanding of how neural circuits control the complex coordinated movement of the tongue. To address this, we need to monitor and quantify its movements, which is challenging. The recent development of micro-X-ray reconstruction of moving morphology (micro-XROMM) provides the critical ability to visualize and quantify movements that typically cannot be seen from the outside with millisecond and micrometer resolution. In this project, Eiman Azim is using micro-XROMM to investigate the mouse brainstem neural circuits that control tongue movements during natural behaviors like chewing and drinking. This work will provide insight into how sensorimotor brain circuits control the rich behavioral repertoire of the tongue and how pathology arises when these functions are disrupted.
Innovation Grant
Plants and animals are constantly interacting with microbes (bacteria, fungi, viruses, etc.). These host-microbe interactions are central to health and disease, growth and development, and ecosystem functioning. Land plants, for example, form close relationships with arbuscular mycorrhiza fungi (AMF), which provide the plants’ root cells with mineral nutrients in exchange for carbon. To balance the costs and benefits of this symbiotic relationship, plants tightly control the fungal colonies through local and systemic signaling pathways, which allows them to coordinate fungal interactions with their current nutrient needs and carbon availability. However, much less is known about how the fungal species regulate their side of this partnership. Lena Mueller is performing dual-species transcriptomics to simultaneously measure plant and fungal gene expression with single-nucleus resolution in a variety of plant-AMF pairings. Her findings will lay the groundwork for understanding mechanisms of microbial control over plants and help identify mechanisms to prevent microbial parasites. This research may also identify strategies to promote beneficial plant-microbe interactions, which could help boost crop yields and reduce the use of chemical fertilizers.
Innovation Grant
Neuroscientists need to be able to image large populations of cells deep within the brain, which is challenging to do with existing microscopy tools. For example, two-photon microscopy allows for imaging of neural activity in deeper layers of the brain, but this relies on a nonlinear excitation process that only lets scientists see the activity in a small volume of tissue. Adam Bowman has identified a method to generate an effective nonlinearity that will allow his lab to get the benefits of two-photon microscopy with standard one-photon excitation. He is now developing time domain optics to allow nanosecond-level control of this microscope illumination. Using these optics, his lab will demonstrate a prototype microscope that can achieve time-gated optical sectioning of brain tissue with orders of magnitude higher throughput than two-photon microscopy and without requiring a spatial pinhole to reject out-of-focus scattered light. He will deploy this technique to improve the readout of fluorescent molecules in the deep brain, which could transform our ability to observe and decode neural activity.
Collaboration Grant
Christian Metallo and Janelle Ayres are investigating a potential topical therapeutic for normalizing microbial dysbiosis to improve skin health. They have discovered sphingolipid species in human skin that they call “very long-chain sphingoid bases” (VLCBs). These are the most abundant sphingolipids produced in the human epidermis, yet their existence and bioactivity have been previously ignored. Preliminary data suggest VLCBs are decreased ~100-fold in patients with atopic dermatitis or eczema, a common inflammatory skin condition marked by altered microbiota. They are now investigating whether VLCB treatment can normalize keratinocyte differentiation and the microbial ecology of the skin in patients with atopic dermatitis. The project leverages their combined expertise in lipidomics, cell biology, and host-microbe interactions to define the functional role of VLCBs in maintaining ecological homeostasis in the skin.
Innovation Grant
Immunotherapies have radically transformed cancer care by using the body’s own immune cells to attack tumors. But there is still a lot of room for improvement, as most tumors lack markers for the immune system to recognize, and promoting more aggressive immune responses as a workaround often results in devastating autoimmune reactions. Salk Assistant Professor Daniel Hollern has an innovative solution to this problem: taking control of the immune system’s B cells and causing them to release anti-tumor antibodies. These antibodies then serve to mark cancerous cells so the immune system can more easily find and attack them. Already, Hollern has found that injecting the antibodies into mice with triple-negative breast cancer can halt tumor growth and even cure the mice. He now plans to expand and develop the concept into an antibody therapy for human cancer patients.
Innovation Grant
Comparing the genomes of different plant species often reveals many genes that have been gained or lost during evolution or domestication. Yet the sheer amount of genetic data prevents scientists from testing evolutionary hypotheses about plants. Now, Salk Research Professor Todd Michael hopes to overcome this challenge by generating plant artificial chromosomes that hold hundreds to thousands of genes. His first step will be to assemble a plant artificial chromosome in yeast; success will mean that researchers can begin developing plant artificial chromosomes in other species. Michael says plant artificial chromosomes will revolutionize scientists’ ability to address fundamental questions about plant evolution—a critical line of inquiry as researchers work to improve crop plant stability and resilience in the face of climate change.
Collaboration Grant
RNA is found in all forms of life and plays many important roles in making proteins and regulating gene expression. Like DNA, RNA is made of building blocks called nucleotides, which are strung together and then folded into a 3D structure. The shape of each RNA molecule then determines its functional capabilities. However, as the molecule accumulates mutations over time, it can change its shape and adopt new functions. Still, little is known about how evolution shapes RNA structure and function. A scientist studying the evolution of an animal species might look at the fossil record and trace how one specimen gradually evolved into another. It is also possible to infer the evolutionary record by comparing the genetic sequences of living organisms, but it is difficult to trace the historical lineage of evolving RNAs. Now, Salk Professor and President Gerald Joyce and Associate Professor Dmitry Lyumkis have proposed a new method for capturing the molecular details of how RNA remodels itself through evolution. They have devised a system to drive and track the evolution of a specific RNA enzyme, collecting the sequence and atomic-resolution structure of each version of the molecule across its entire lineage. Their findings will help explain how advantageous RNA structures emerge and are stabilized during evolution. More tangibly, the project will produce a generalizable workflow for studying RNA structure and a rich dataset to support the use of RNAs in biology, medicine, and biotechnology.
Innovation Grant
The human brain is organized by functional regions, with each region containing cell types with specialized functions. For example, astrocytes—a neuron-supporting brain cell—may look and act differently in one region of the brain compared to another. Despite this variability in astrocyte subtypes, a tool does not yet exist to parse through the differences between them—Salk Associate Professor Nicola Allen wants to change that. Beginning with the brain regions responsible for vision, Allen will develop tools to manipulate astrocytes in specific sub-regions using CellREADR, then assess how those manipulations impact visual function, in turn learning about the region-specific role of astrocytes in the visual system. If successful, the tool can be applied to study other subclasses of astrocytes around the brain and, in turn, help scientists better understand brain function.
Innovation Grant
Killing pathogens is not the body’s only defense against infection. “Cooperative defenses” are an entirely separate defense strategy that allows the body to work with a pathogen, rather than against it. The system ameliorates physiological symptoms of infection with either neutral or beneficial impact on the infectious pathogen causing those symptoms. Salk Professor Janelle Ayres is leading the charge on characterizing the mechanisms of cooperative defenses for infection treatment and determining how their discoveries can be applied to non-infectious diseases. Ayres, alongside postdoctoral researcher Katia Troha, discovered one cooperative defense mechanism that enhances kidney function—findings they hope can lead to strategies for kidney regeneration. By probing this kidney mechanism further and expanding the research to cover other areas of the body, Ayres’ investigation will show how the cooperative defense system helps animals optimize organ function and rebuild tissues in disease states.
Innovation Grant
Most of the DNA in your cells consists of instructions that dictate when and where genes should be active. Uncovering which parts of the instructions are read by each of the many different cell types is key to understanding cell function, dysfunction, and diversity. Professor Edward Callaway and Assistant Professor Pallav Kosuri will create a new screening platform based on cutting-edge MERFISH technology to identify pieces of DNA called enhancers, which regulate gene expression in different cell types. By bringing down the cost of these screening tests by a factor of a hundred compared to current methods, the MERFISH-based platform will make it possible to explore gene control in thousands of different kinds of cells at once.
Innovation Grant
From a buzzing beehive to a bustling school campus, social colonies are key to the evolutionary success of animals. What biologically creates and sustains these social colonies, however, is yet to be fully understood. Investigation into the genes that control social behaviors in animals has long been limited by the lack of an effective model organism. But Associate Professor Kenta Asahina thinks the sweat bee is the solution. Using a thorough understanding of fruit fly genetics as a springboard, he hopes to establish sweat bees as a novel model organism for exploring the evolutionary origins and brain basis of social behaviors.
Innovation Grant
Neuroscientists are working to unravel how the brain controls movement. Current approaches to track movement ignore the body’s biomechanics and other constraints, like gravity. Associate Professor Eiman Azim and Salk Fellow Talmo Pereira will create a computational model of how the brain produces behavior that takes into account what we know about the nervous system and structure of the body—brains, bones, and muscles. They hope this new approach opens doors for neuroscientists to study the relationship between the brain, body, and movement with higher accuracy than ever before.
Innovation Grant
Within the nucleus of the cell, DNA is wrapped around specialized packaging proteins called histones to form chromatin fibers. Within chromatin, both the DNA and histone components can be modified by the attachment of small chemical tags to affect the level of chromatin compaction and the expression of underlying genes. In addition, some of these modifications can alter the 3D organization of chromatin within the nucleus. To better understand the connections between chromatin modifications, genome organization, and gene regulation in plants, Associate Professor Julie Law and Assistant Professor Jesse Dixon will use a genomic analysis technique called Hi-C to explore how genetically altered chromatin states impact genome organization.
Pilot Award from the Innovation Program and Salk Cancer Center
Chromosomes have little endcaps called telomeres, which shorten with age. As they shorten, the process of replicating the chromosomes becomes more difficult—occasionally causing what’s known as replicative crisis. Once in a state of replicative crisis, a cell either dies or becomes cancerous. In cases where the cell dies, short telomeres send distress signals to the cell’s mitochondria to activate another signaling pathway that promotes cell death. Using the latest imaging technologies, Salk Associate Professor Dmitry Lyumkis, Professor Gerald Shadel, Professor and CSO Jan Karlseder, and Scripps Assistant Professor Danielle Grotjahn are embarking on an effort to chart the relationship between telomeres, mitochondria, and cell death, in turn uncovering how mitochondrial form and function impact age-related cancer development.
Collaboration Grant
Scientists are using artificial intelligence (AI) to assist with the clinical diagnosis of cancer through video-based imaging, offering a non-invasive option for predicting the trajectory of cancer progression from body language. Professor Christian Metallo, Assistant Professor Dannielle Engle, and Salk Fellow Talmo Pereira believe the use of AI can greatly improve how we measure the health status of animals receiving cancer treatment. The team will build AI-based models to relate changes in mouse behavior and diet to pancreatic cancer progression and whole-body physiology. Their findings will further AI application in cancer treatment while improving our scientific understanding of diet and chemotherapy, key factors important to all cancer patients.
Collaboration Grant
Inflammation is characteristic of a wide variety of diseases, including Alzheimer’s, and plays a role in neurodegeneration and aging. Salk’s Professor Rusty Gage, Professor Christian Metallo, and Associate Professor Axel Nimmerjahn want to see how specialized brain-resident immune cells called microglia may cause or perpetuate inflammation, or if they could be therapeutic targets for reducing inflammation. To investigate the role of microglia in inflammation, they plan to create a first-of-its-kind human brain organoid (a three-dimensional collection of cells that mimics features of human tissues) populated with microglia, which they can use to characterize and observe microglia in a human brain-like setting.
Innovation Grant
Not everything is written in our genes—after genes are translated into proteins, cells still modify those proteins to regulate function. Salk Professor Tony Hunter discovered that cells sometimes add a phosphate group—a chemical tag—to a protein component called tyrosine. This event, tyrosine phosphorylation, drives critical cell communications and often malfunctions in cancer. Now it’s emerging that phosphorylated histidine (pHis), another protein component, may play similar roles. Hunter’s lab generated the first antibodies recognizing pHis. Now they will engineer antibodies for higher binding affinity towards pHis proteins and use them to probe pHis function in health and disease.
Innovation Grant
It’s long been accepted that a single neuron can send only one type of signal, but recent studies have shown that single neurons can store two different pools of transmitters. Assistant Professor Sung Han will explore how single neurons orchestrate the release of different transmitters—glutamate vs. neuropeptides, for example. With this idea, he and his lab are currently investigating the neuronal coding logic of transmitter co-transmission to encode various information. These studies will transform the field’s current understanding of neuronal communication.
Innovation Grant
Plants use sensory proteins to detect and respond to touch signals from animals and neighboring plants. Professor Joanne Chory, Associate Professor Sreekanth Chalasani, and Staff Scientist Carl Procko will investigate these proteins to see if they are sensitive to high-frequency sound waves. The work could help scientists modify how plants behave in the presence of other plants and contribute to the growing body of research on “sonogenetics,” a method Chalasani developed for non-invasively controlling cells with sound waves. Their findings could also lead to new ways for treating conditions like chronic pain, epilepsy and PTSD.
Innovation Grant
The ease with which researchers can track body parts in motion, such as hands and feet, has improved in recent years due to advances in machine learning and computer vision. However, these methods still require researchers to manually label each body part, which is time-consuming and increasingly impractical as the number of labels on the body grows. Assistant Professor Eiman Azim is now developing automated approaches to simultaneously track hundreds to thousands of points on the body. These methods will provide better ways to examine how brains control movement and insights into how neurodegenerative disease and injury disrupt behavior.
Collaboration Grant
Salk Professor Janelle Ayres recently discovered that uninfected animals respond differently to an infection depending on whether they had previously interacted with asymptomatic or symptomatic infected group mates. Ayres will now collaborate with Salk Professors Joseph Noel and Christian Metallo to determine if asymptomatic and symptomatic individuals emit different chemicals—signals that help group mates respond to the infection. The new team will combine expertise in animal physiology, evolutionary theory and infectious diseases with metabolic biochemistry and mass spectrometry. The methods they develop may help diagnose community spread of infectious diseases and develop new treatments based on emitted chemicals.
Innovation Grant
The Chalasani lab is seeking to understand how animals make decisions. Associate Professor Sreekanth Chalasani and graduate student Jess Haley will use a novel microscopy system to record the activity of most, if not all, neurons in the C. elegans “brain” as it learns, remembers and makes decisions. They will discover neuronal changes associated with each of these three phenomena (learning, memory and decision-making), providing a framework for analyzing more complex brains.
Innovation Grant
RNA Polymerase III (Pol III) mutations cause hypomyelinating leukodystrophy (HMLD), a fatal neurodegenerative disease caused by defective CNS nerve myelination. In yeast, conserved Pol III HMLD disease mutations cause growth defects, which are rescued by inhibiting the sumoylation pathway. Professor Tony Hunter’s lab will investigate if Pol III mutant-derived neurodegenerative phenotypes are recapitulated in human neural cells and in animal models, and, if so, how the sumoylation pathway contributes to such phenotypes. This would implicate sumoylation as a target for rescue of Pol III-related neurodegenerative diseases.
Innovation Grant
Intercellular interactions activate signaling pathways and cause phenotypic changes. The Wahl lab suspects such interactions contribute to cancer metastasis, a deadly killer of cancer patients, yet there are no methods to track the constellation of cells that a cancer cell interacts with in the metastatic niche. Professor Geoffrey Wahl, postdoctoral researcher Nikki Lytle and project scientist Leo Li will develop Contact Tracing, an innovative tool to indelibly label interacting cells for subsequent identification, isolation and analysis. This may reveal cellular relationships that drive metastatic progression.
Collaboration Grant
Aging is associated with dysfunctional immune responses, but no one knows the initiating events behind these processes. Professor Susan Kaech, Associate Professors Diana Hargreaves and Ye Zheng, and Assistant Professor Jesse Dixon believe epigenetic influences are particularly vulnerable to age and that many of the age-related changes in immune function and inflammation stem from epigenetic dysfunction. They will determine the effects of age on these factors during a viral infection, identify key factors in aging-associated decline in immunity, and facilitate efforts to improve immune response in older patients.
Collaboration Grant
ALS is hallmarked with delayed adult onsets at unpredictable sites followed by devastating and progressive spread. Professors Sam Pfaff and Axel Nimmerjahn, Associate Professor Nicola Allen, and Assistant Professor Eiman Azim believe that disease onset is accelerated or triggered by secondary environmental insults. An understanding of these environmental interactions could lead to avenues that would delay ALS onset so they are investigating how genetic models of ALS react to these environmental insults, which will begin to define how genetic predisposition and secondary environmental events converge to trigger ALS.
Collaboration Grant
Professors John Reynolds, Juan Carlos Izpisua Belmonte, and Rusty Gage will examine the hallmarks of aging in animal models to determine whether mobile DNA elements, called LINE1 retrotransposons, can be manipulated to slow or reverse aging to create an innovative healthy aging intervention.
Collaboration Grant
Professor Martyn Goulding, Associate Professor Axel Nimmerjahn, and Assistant Professor Sung Han will investigate how sensory signals from the skin, the biggest sensory organ, are processed as they travel to the brain. The project will potentially reveal new targets for sensory dysfunction, which can occur with chronic pain and autism spectrum disorders.
Innovation Grant
One of the holy grails of circadian biology research is to understand what determines whether an animal is active during the day (diurnal) or active during the night (nocturnal). To begin to answer this question, Professor Satchidananda Panda will measure changes in hormones and gene activity as two species of animals— the owl monkey and mouse—switch between diurnal and nocturnal lifestyles. One potential outcome of this work will be strategies for improving the health and life quality of shift workers, a growing fraction of the worldwide workforce.
Innovation Grant

Edward Stites
The RAS protein is frequently mutated in some of the most difficult to treat cancers, including lung and colon. To better understand the contribution of RAS to cancer, Assistant Professor Edward Stites will activate a particularly deadly version of RAS in the microscopic worm C. elegans, a widely used model organism. Subsequent genetic and chemical screens will help to reveal new drugs and therapeutic strategies for treating RAS-associated cancers.
Innovation Grant
Cancer cells are metabolically greedy, which often leads to nutrient depletion within and around a tumor. Professor and Director of the NOMIS Center for Immunobiology and Microbial Pathogenesis Susan Kaech hypothesizes that this lack of nutrients starves immune cells that might otherwise recognize and eliminate the tumor. The team will map the nutrient landscape of different tumors, with the goal of identifying specific nutrients (metabolites) that boost immune cell effectiveness. Results will help to improve current anti-cancer immunotherapies.
Rose Hills Foundation Innovation Grant
Assistant Professor Dannielle Engle was named The Rose Hills Foundation’s 2020-2021 Innovator Grant Program awardee. The award provides $100,000 for Engle to investigate how the sugar CA19-9 makes pancreatic cancer more aggressive, increases metastatic spread, and interacts with metastatic sites. As most pancreatic cancer patients are diagnosed with metastatic disease, blocking CA19-9 interactions may intercept metastatic spread.
Rose Hills Foundation Innovation Grant
Assistant Professor Kenta Asahina was named The Rose Hills Foundation’s 2019-2020 Innovator Grant Program awardee, in coordination with Salk’s Innovation Grants Program. The award provides $100,000 for Asahina to explore a novel approach to studying aging. Asahina, who is holder of The Helen McLoraine Developmental Chair in Neurobiology, will study how the nervous system can contribute to lifespan difference between sexes. Females in many animal species, including humans, live longer than males. Asahina’s work aims to provide a novel paradigm for studying aging, with a potential to understand the biological mechanisms that specify human lifespan.
Collaboration Grant
Professors Ronald Evans and Susan Kaech and Associate Professor Ye Zheng, will lead a team in exploring if a healthy diet and exercise reduces levels of inflammation and renders tumor cells more sensitive to the immune system, with the goal of expanding the efficacy of immunotherapies.
Collaboration Grant
Professors Alan Saghatelian, Joseph Noel, and Jan Karlseder will undertake a multi-pronged approach to develop small-molecule inhibitors of the DNA repair regulator CYREN, with the goal of specifically sensitizing tumor cells to genotoxic therapy.
Innovation Grant
Thomas Albright, professor and director of the Vision Research Laboratory, will work with Staff Scientist Sergei Gepshtein to investigate the neurological basis of how individuals recognize others, which could lead to better ways to identify suspects during criminal investigations. The goal is to help reduce cases where innocent people are misidentified during lineups.
Innovation Grant
Wolfgang Busch, associate professor, Uri Manor (core director, Waitt Advanced Biophotonics Core), and Saket Navlakha (assistant professor) will explore the biological algorithms that guide how plants grow and pattern their root systems in search of nutrients. This research may uncover how plants can efficiently find water and other elements in the soil, advancing Salk’s efforts to engineer plants capable of surviving increasingly erratic climate patterns.
Innovation Grant
Jesse Dixon, a Helmsley-Salk Fellow, is exploring how mutations in individual cells can lead to the development of cancer. His team seeks to understand tumors’ evolutionary histories and potentially reveal new strategies that can halt tumor progression by interrupting the evolution of cells from normal to cancerous.
Innovation Grant
Susan Kaech, professor and director of the NOMIS Center for Immunobiology and Microbial Pathogenesis, and Ronald Evans, professor and director of the Gene Expression Laboratory and a Howard Hughes Medical Institute investigator, will embark on a study of lipid metabolism as a weapon in the fight against pancreatic cancer, a notoriously difficult-to-treat disease.
Innovation Grant

Juan Carlos Izpisua Belmonte
Professor Juan Carlos Izpisua-Belmonte of the Gene Expression Laboratory seeks to determine whether trans-generational epigenetic inheritance can take place in mammals. If possible, this would mean that the experiences that have shaped the genetic expression of parents (e.g., adaptations to environmental challenges) could be passed to children—a significant question in evolutionary biology which remains unanswered.
Innovation Grant
Associate Professor Sreekanth Chalasani of the Molecular Neurobiology Laboratory alongside Research Associate Chen-Min Yeh and Staff Scientist Gerald Pao seek to answer the question of whether or not brain activity can be used to control a robot. They will leverage advanced live microscopy techniques, in addition to supercomputer technology, to see whether or not the brain activity of zebrafish larvae can control a fish robot.
Innovation Grant
Professor and Laboratory Head David Schubert of the Cellular Neurobiology Laboratory will work with Staff Scientist Antonio Currais to identify new drug candidates for Alzheimer’s disease using screens for mitochondrial dysfunction. Specifically, they will look at a large library of plant extracts that have pharmacological value to see any have protective traits that are able to preserve mitochondrial function—one of the earliest clinical challenges in Alzheimer’s.
Innovation Grant
Joseph Ecker, a Professor in the Plant Molecular and Cellular Biology Laboratory and the Director of the Genomic Analysis Laboratory, is working to develop a method that allows researchers to record the transcriptional activity within a cell into the genetic code so that they can analyze the cascade of transcriptional events that occur during an organism’s development as well as cell reprogramming.
Innovation Grant
Professor Edward Callaway in the Systems Neurobiology Laboratory is undertaking a project that will develop innovative methods for flexible, high-throughput analysis of specific brain-cell types across any species, including humans, that can identify the genetic enhancers that restrict expression of genes that have been passed from one cell (or whole organism) to another.
Rose Hills Foundation Innovation Grant

Xin Jin
Xin Jin, an associate professor, will explore how networks of neurons communicate with one another to relay messages from the brain to the limbs. His lab uses research techniques in novel combinations to discover previously unknown connections between brain regions and how they contribute to movement control. Jin’s work may lead to new avenues for treating disorders such as Parkinson’s disease.












































