
August 5, 2009
LA JOLLA, CA—Rather than sticking to a single DNA script, human brain cells harbor astonishing genomic variability, according to scientists at the Salk Institute for Biological Studies. The findings, to be published in the Aug. 5, 2009, advance online edition of Nature, could help explain brain development and individuality, as well as lead to a better understanding of neurological disease.
The team, led by Fred Gage, Ph.D., a professor in the Salk’s Laboratory of Genetics and holder of the Vi and John Adler Chair for Research on Age-Related Neurodegenerative Diseases, found that human brain cells contain an unexpected number of so-called mobile elements—extraordinary pieces of DNA that insert extra copies of themselves throughout the genome using a “copy and paste” mechanism.
“Jumping genes” could go a long way towards explaining brain development and individuality.
Image: Courtesy of Jamie Simon, Salk Institute for Biological Studies.
“This is a potential mechanism to create the neural diversity that makes each person unique,” says Gage. “The brain has 100 billion neurons with 100 trillion connections, but mobile pieces of DNA could give individual neurons a slightly different capacity from each other.”
The only other human cells known to remodel their genome are the cells of the immune system. There the genes coding for antibodies are shuffled to create the necessary variety of antibodies capable of recognizing an infinite number of distinct antigens.
In earlier work, Gage had already shown that mobile pieces of DNA known as LINE-1 elements (short for Long interspersed element 1) randomly add extra copies to the genome of mouse brain cells. But whether or not the same process, colloquially referred to as “jumping,” held true for neurons in human brains had been a matter of some debate.
“It is known that these mobile elements are important in lower organisms, such as plants and yeast, but in mammals they are generally considered to be remnants of our past,” says Gage. “Yet they are extremely abundant. Approximately 50% of the total human genome is made up of remnants of mobile elements. If this were true junk, we would be getting rid of it.”
Initially, the Salk researchers simply tried to identify jumping events in human brain cells grown in a dish, as they had with mice. They succeeded, but neither Gage nor postdoctoral researcher and first author Nicole Coufal, Ph.D., was satisfied. They wanted to know whether this phenomenon occurred in people as well.
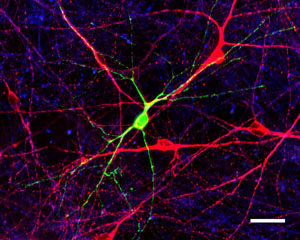
During the early stages of human nervous system development LINE-1 elements become active (indicated by green flourescence) possibly affecting neuronal function.
Image: Courtesy of Dr. Nicole Coufal, Salk Institute for Biological Studies.
In mice, gene jumping is limited to the brain and to germ cells, the reproductive cells in our bodies, since these kinds of rapid changes would be detrimental in essential organs such as the heart or kidneys. If LINE-1 elements were indeed on the move in humans, there should be more copies in the brain than in the heart or liver, where Gage and Coufal did not expect LINE-1 jumping to occur.
When Coufal measured matched samples (brain versus other body tissues) from numerous individuals, she found that some brain samples had as many as 100 extra copies per cell. “This was proof that these elements really are jumping in neurons,” explains Coufal. Strikingly, it also means that not all cells are created equal—humans are true chimeras since the DNA in their brain cells is different from the DNA in the rest of their cells.
In this way, mobile elements may actually drive evolution, creating more diversity than would occur through normal cell division (which makes an exact copy of the genome, save the occasional typo). “It’s a different way of looking at diversity,” says Gage. “The brain lives for 80 years with the environment coming at us unpredictably, and this provides an added element of adaptability. It makes sense that there would be this added level of complexity.”
Trying to explain why only brain cells harbor truly “mobile” elements, Coufal looked at the LINE-1 promoter, which is the switch that turns LINE-1 elements on and off, and she discovered that in the brain, the switch is generally on, whereas in other tissues it is permanently locked in the “off” position.
More importantly, knowing that LINE-1 elements are jumping around in the brain opens up a new way to look at neurological disorders. Explains Coufal, “Dysregulated jumping could be contributing to the problems seen in these conditions.” The Salk researchers are planning to look for changes in jumping frequency in the brains of individuals affected by Rhett’s syndrome and other diseases.
In addition to Drs. Coufal and Gage, researchers contributing to this study include Grace Peng, Gene Yeo, Ph.D., Yangling Mu, Ph.D., and Michael Lovci, Ph.D. (now at the University of California, San Diego), José Garcia-Perez, Maria Morell and K. Sue O’Shea at the University of Michigan Medical School, and John V. Moran at the University of Michigan Medical School and Howard Hughes Medical Institute.
For information on the commercialization of this technology, please contatct the Salk Office of Technology Development at (858) 453-4100, Ext. 1278.
About the Salk Institute for Biological Studies: The Salk Institute for Biological Studies is one of the world’s preeminent basic research institutions, where internationally renowned faculty probe fundamental life science questions in a unique, collaborative, and creative environment. Focused both on discovery and on mentoring future generations of researchers, Salk scientists make groundbreaking contributions to our understanding of cancer, aging, Alzheimer’s, diabetes, and cardiovascular disorders by studying neuroscience, genetics, cell and plant biology, and related disciplines.
Faculty achievements have been recognized with numerous honors, including Nobel Prizes and memberships in the National Academy of Sciences. Founded in 1960 by polio vaccine pioneer Jonas Salk, M.D., the Institute is an independent nonprofit organization and architectural landmark.
– Claire Attwooll
Office of Communications
Tel: (858) 453-4100
press@salk.edu