Associate Professor
Plant Molecular and Cellular Biology Laboratory

In any one organism, every cell has nearly identical genetic information, yet not all cells look or act the same. One cause of this amazing diversity is the presence of chemical tags, called epigenetic modifications, which decorate both the DNA and packaging proteins that organize the DNA within the nucleus. The patterns of these chemical tags differ in each cell type and help instruct the function of cells by indicating which genes should be turned on and which should be ignored. Knowing the effect of these epigenetic changes on a cell’s behavior can help us to understand health and disease, but manipulating epigenetics in mammalian cells is often lethal, making these changes hard to study. Fortunately, similar manipulations are viable in a plant model, making this an excellent system to understand the roles of epigenetic modifications.

Rather than using animal cells to study epigenetic modifications, Julie Law is turning to the small flowering plant Arabidopsis thaliana. Unlike mammals, Arabidopsis thaliana plants are more tolerant of changes to their epigenome, making it easier to study the effects of altering these chemical tags. Using this plant, Law is studying how epigenetic modifications are recognized and translated into the desired response by the cell. In particular, she focuses on characterizing several newly identified families of proteins involved with DNA packaging and gene expression, called chromatin binding proteins. By employing genetic, biochemical and genomics approaches, Law aims to not only determine the epigenetic marks recognized by these protein families, but also to identify their interacting partners and their effects on gene expression. Although Law’s research utilizes a plant model, her findings will also hold lessons for human biology as many of the genes involved in adding or removing epigenetic marks are the same in plants as in mammals. Ultimately, her research paves the way for understanding the role of the epigenome in both agriculture and human health.
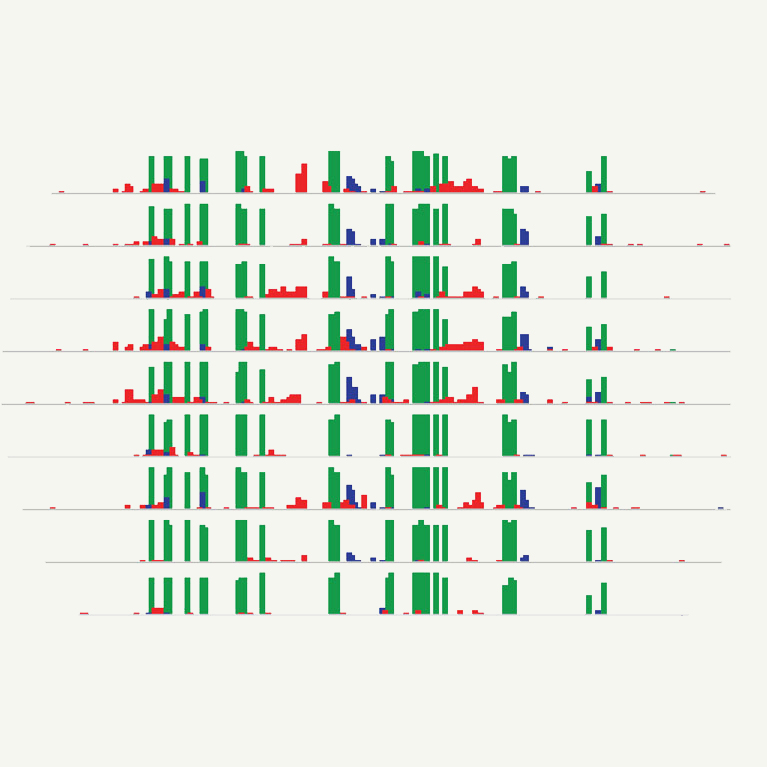
With colleagues, Law provided mechanistic insights into the targeting of two specialized RNA polymerases (Pol-IV and Pol-V) in the Arabidopsis thaliana genome. These findings provide insight into how epigenetic modifications might be targeted to specific genes for crop improvements or therapeutic benefits to human health.

Taking a biochemical approach, Law used proteins with known roles in a process called DNA methylation to identify a handful of additional proteins never before linked to epigenetic modifications. Knowledge of the new proteins enhances our understanding of the processes influencing a cell’s epigenome.
Law and her colleagues also revealed, at the level of individual atoms, the precise regions of several proteins that are critical in recognizing specific epigenetic modifications. These studies provide a detailed view of how these proteins function and could reveal how their mutation can lead to epigenetic changes that manifest as developmental defects or the progression of diseases, such as cancer.
BS, Biochemistry and Biophysics, Oregon State University
PhD, Biochemistry, Johns Hopkins University School of Medicine
Postdoctoral Fellow, University of California, Los Angeles