
March 30, 2011
LA JOLLA, CA—The moment we open our eyes, we perceive the world with apparent ease. But the question of how neurons in the retina encode what we “see” has been a tricky one. A key obstacle to understanding how our brain functions is that its components—neurons—respond in highly nonlinear ways to complex stimuli, making stimulus-response relationships extremely difficult to discern.
Now a team of physicists at the Salk Institute for Biological Studies has developed a general mathematical framework that makes optimal use of limited measurements, bringing them a step closer to deciphering the “language of the brain.” The approach, described in the current issue of the Public Library of Science, Computational Biology, reveals for the first time that only information about pairs of temporal stimulus patterns is relayed to the brain.
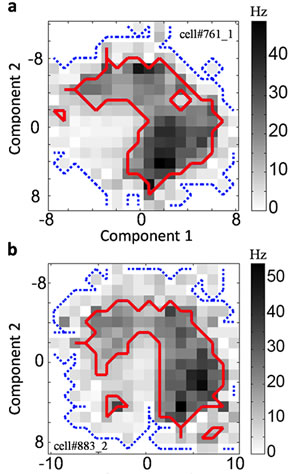
Spike distributions for neurons responding to two features can have shapes that are difficult to understand.
Image: Courtesy of Dr. Tatyana Sharpee, Salk Institute for Biological Studies
“We were surprised to find that higher-order stimulus combinations were not encoded, because they are so prevalent in our natural environment,” says the study’s leader Tatyana Sharpee, Ph.D., an assistant professor in the Computational Neurobiology Laboratory and holder of the Helen McLorraine Developmental Chair in Neurobiology. “Humans are quite sensitive to changes in higher-order combinations of spatial patterns. We found it not to be the case for temporal patterns. This highlights a fundamental difference in the spatial and temporal aspects of visual encoding.”
The human face is a perfect example of a higher-order combination of spatial patterns. All components—eyes, nose, mouth—have very specific spatial relationships with each other, and not even Picasso, in his Cubist period, could throw the rules completely overboard.
Our eyes take in the visual environment and transmit information about individual components, such as color, position, shape, motion and brightness to the brain. Individual neurons in the retina get excited by certain features and respond with an electrical signal, or spike, that is passed on to visual centers in the brain, where information sent by neurons with different preferences is assembled and processed.
For simple sensory events—like turning on a light, for example—the brightness correlates well with the spike probability in a luminance-sensitive cell in the retina. “However, over the last decade or so, it has become apparent that neurons actually encode information about several features at the same time,” says graduate student and first author Jeffrey D. Fitzgerald.
Example of the flickering light stimulus presented during the experiment.
Movie: Courtesy of Dr. Tatyana Sharpee, Salk Institute for Biological Studies
“Up to this point, most of the work has been focused on identifying the features the cell responds to,” he says. “The question of what kind of information about these features the cell is encoding had been ignored. The direct measurements of stimulus-response relationships often yielded weird shapes [see Figure 1, for example], and people didn’t have a mathematical framework for analyzing it.”
To overcome those limitations, Fitzgerald and colleagues developed a so-called minimal model of the nonlinear relationships of information processing systems by maximizing a quantity that is referred to as noise entropy. The latter describes the uncertainty about a neuron’s probability to spike in response to a stimulus.
When Fitzgerald applied this approach to recordings of visual neurons probed with flickering movies, which co-author Lawrence Sincich and Jonathan Horton at the University of California, San Francisco, had made, he discovered that on average, first-order correlations accounted for 78 percent of the encoded information, while second-order correlations accounted for more than 92 percent. Thus, the brain received very little information about correlations that were higher than second order.
“Biological systems across all scales, from molecules to ecosystems, can all be considered information processors that detect important events in their environment and transform them into actionable information,” says Sharpee. “We therefore hope that this way of ‘focusing’ the data by identifying maximally informative, critical stimulus-response relationships will be useful in other areas of systems biology.”
The work was funded in part by the National Institutes of Health, the Searle Scholar Program, The Alfred P. Sloan Fellowship, the W.M. Keck Research Excellence Award and the Ray Thomas Edwards Career Development Award in Biomedical Sciences.
About the Salk Institute for Biological Studies:
The Salk Institute for Biological Studies is one of the world’s preeminent basic research institutions, where internationally renowned faculty probe fundamental life science questions in a unique, collaborative and creative environment. Focused both on discovery and on mentoring future generations of researchers, Salk scientists make groundbreaking contributions to our understanding of cancer, aging, Alzheimer’s, diabetes and infectious diseases by studying neuroscience, genetics, cell and plant biology and related disciplines.
Faculty achievements have been recognized with numerous honors, including Nobel Prizes and memberships in the National Academy of Sciences. Founded in 1960 by polio vaccine pioneer Jonas Salk, M.D., the Institute is an independent nonprofit organization and architectural landmark.
Office of Communications
Tel: (858) 453-4100
press@salk.edu