
February 17, 2016
Salk Institute and collaborators show key molecular differences between viruses as they interact with host DNA
Salk Institute and collaborators show key molecular differences between viruses as they interact with host DNA
LA JOLLA—Using cutting-edge imaging technology, Salk Institute and Harvard Medical School researchers have determined the structure of a protein complex that lets viruses similar to the human immunodeficiency virus (HIV) establish permanent infections within their hosts.
Contrary to previous assumptions, the newly detailed viral protein complex structure indicates that this type of molecular architecture differs across retroviruses. This information helps reveal how retroviruses insert their genomic information into human cells and may have implications not only for treating diseases like HIV, but also for improving gene therapy methods to deliver new DNA to patients with genetic mutations.
“The details of how retroviruses integrate differ far more than previously thought and lead to entirely distinct patterns of infection,” says Dmitry Lyumkis, a 索尔克研究员 and co-senior author of the new paper, published in 自然 on February 18, 2016.
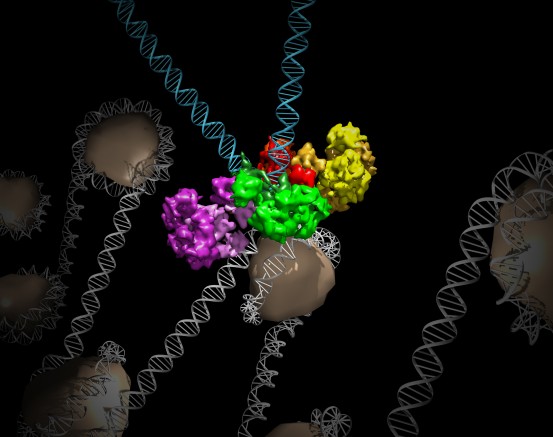
点击此处 用于高分辨率图像。.
版权:萨克研究所
A retrovirus inserts its genetic information into the host’s own genome, essentially turning the host’s cells into virus-making factories. In the case of the well-known retrovirus HIV, those viral genes integrate into human immune cells, eventually killing them. A long-standing mystery—caused in part by the lack of knowledge about the protein complex structure—has been what dictates where viral DNA is inserted in the human genome.
A protein complex called an intasome is responsible for irreversibly inserting viral DNA into human DNA, a vital step in allowing retroviruses to cause disease. Since the intasome of HIV itself is tricky to study, most of what’s been known about intasomes is based on work from another retrovirus called prototype foamy virus, or PFV. In light of specimen-related challenges pertaining to HIV intasomes, Lyumkis and Engelman, along with their colleague Peter Cherepanov at the Francis Crick Institute, set out to determine the structure of the intasome from mouse mammary tumor virus (MMTV), which in some ways is more closely related to HIV than PFV.
“The MMTV intasome structure defines an unexpected novel paradigm for the structural basis of retroviral DNA integration,” says co-senior author Alan Engelman, professor of medicine at the Dana-Farber Cancer Institute and Harvard Medical School.
To determine MMTV’s molecular structure, the team used an up-and-coming technique called cryo-electron microscopy (cryoEM), an approach that has several advantages over traditional imaging techniques. For example, researchers don’t need to coax proteins into crystal forms to image them, a hit-or-miss process that is required for the more common technique of x-ray crystallography. In cryoEM, proteins are instead directly frozen in a liquid solution. Then, by measuring how beams of electrons are deflected off of the frozen samples, scientists can determine the structure of the proteins.
Lyumkis used this cutting-edge technology to detail the structure of MMTV intasomes bound to strands of viral DNA (the way the structure would look as it invaded a host cell). The team found that while the PFV complex is composed of four protein components, called integrases, bound to two strands of viral DNA, the MMTV assembly has eight integrase molecules per two pieces of viral DNA.
This key structural difference, Lyumkis says, means that the complexes interact differently with the host DNA when they are poised to insert viral DNA into the genome. PFV prefers spots in the host genome that are highly bent, while MMTV chooses straight stretches of DNA. The preference for HIV remains a mystery, but the new observations provide an unexpected molecular framework within which to interpret past and future data on HIV and other retroviruses.
The teams of Lyumkis and Engelman are now working to understand the series of events that the MMTV intasome complex goes through during the steps of viral integration, from binding viral DNA to capturing host DNA and catalyzing the insertion of viral DNA into the host’s genome. Lyumkis also plans to apply cryoEM techniques to study the HIV intasome molecular complex itself. “The technological infrastructure is in place so now I think we can tackle these tricky and poorly behaved specimens using cryoEM,” adds Lyumkis.
Other researchers on the study were Allison Ballandras-Colas and Tamaria G. Dewdney of the Dana-Farber Cancer Institute and Harvard Medical School; Monica Brown of the Salk Institute; Nicola J. Cook and Peter Cherepanov of the Francis Crick Institute; and Borries Demeler of the University of Texas Health Sciences Center.
这项工作以及参与研究的学者得到了以下机构资助: 美国国立卫生研究院, , 那个 Leona M. 和 Harry B. Helmsley 慈善信托基金, , 那个 国家科学基金会 and the San Antonio Cancer Institute.
日记
自然
作者
Allison Ballandras-Colas, Tamaria G. Dewdney and Alan Engelman of Harvard Medical School; Monica Brown and Dmitry Lyumkis of the Salk Institute; Nicola J. Cook and Peter Cherepanov of the Francis Crick Institute; and Borries Demeler of the University of Texas Health Sciences Center
宣传办公室
电话:(858) 453-4100
press@salk.edu
萨尔克研究所是一个独立的非营利性研究机构,由首个安全有效的脊髓灰质炎疫苗的研发者乔纳斯·索尔克于1960年创立。该研究所的使命是推动以合作、敢于冒险为特点的基础性研究,以应对癌症、阿尔茨海默病和农业脆弱性等社会最紧迫的挑战。这项基础科学支撑着所有的转化研究,产生有助于全球新药和创新的见解。.