
2026 年 2 月 17 日
Salk Institute scientists show how proteins regulate our genome’s 3D shape to influence gene expression and cell identity
Salk Institute scientists show how proteins regulate our genome’s 3D shape to influence gene expression and cell identity
洛杉矶-我们的 DNA 是如何储存造就人类所需的大量信息的?如果储存错误会发生什么? 杰西·迪克森,医学博士,哲学博士, 多年来,他一直在研究这个基因组在三维空间中的折叠方式——他知道,如果折叠不正常,可能导致癌症和发育障碍,包括与自闭症相关的障碍。他实验室的最新研究表明,人们越来越认识到基因组的三维组织是不断变化的。通过使用不同类型的人类细胞,他的实验室表明,这种动态的基因组展开和重新折叠过程在基因组的不同部分以不同的速率发生,这反过来又影响基因的调控和表达。.
The study, published in Nature Genetics on February 16, 2026, and funded by both federal research grants and private philanthropy, may point to targets for blocking the dysfunctional folding that leads to cancers and developmental disorders.
“There are six billion base pairs in your genome, and in the last decade we’ve been learning about the molecular machines that fold and organize that massive amount of information,” says Dixon, senior author of the study and associate professor and holder of the Helen McLoraine Developmental Chair at Salk. “What’s interesting is that this folding doesn’t just happen once and then the genome stays put—it seems to be constantly unfolding and refolding. Our study gives us a better idea of where and how often the genome is doing this, which ultimately adds to our understanding of those molecular machines, and, in turn, what may be going on when they dysfunction during cancers or developmental disorders.”
Each human cell contains two meters of DNA—critical code that brings every protein, structure, and cellular process to life. Within that DNA code are tens of thousands of genes, which are short stretches of code that can be used to regulate or produce proteins.
This crucial information must be stored and organized in such a way that it can fit inside a cell’s nucleus 和 move around to change gene accessibility 和 be strategically maneuvered to bring together regions that need to interact but that are relatively far apart. Cells have cleverly found a way to knock out all three of these needs at once: loops! Loops are tightly mediated by a protein complex called cohesin, which works alongside an accessory protein, NIPBL, that helps cohesin move along DNA to form these loops.
Recent studies have shown that these cohesin-mediated loops are constantly forming and disassembling. This new understanding of genome folding as a dynamic process inspired a host of new questions: How often is DNA looping and un-looping? Is every part of the genome equally dynamic? What role is NIPBL playing in this movement?
“Current data around the spatial organization of the genome suggest that genome folding has little impact on gene expression—but we thought, perhaps we just aren’t looking at it in the right way,” says first author Tessa Popay, PhD, a postdoctoral researcher in Dixon’s lab. “By specifically disrupting folding dynamics, we were able to identify the aspects of spatial genome organization that contribute to gene regulation and expression.”
The Salk team first depleted NIPBL in immortalized human retinal pigment epithelial (RPE-1) cells to see what impact that would have on loop dynamics. Without NIPBL, cohesin could no longer efficiently move along DNA and form loops. Unable to create new loops, the genomes unfolded—but not uniformly. Rather, some regions of the genome unfolded relatively quickly, while others did so over the course of many hours.
Curiously, the relative stability of different genome areas appeared to be related to functional differences. Loops that were forming and unraveling over many hours were associated with silent genome regions—stretches of the DNA where the genes weren’t in use. Loops that turned over more quickly were associated with expressed genome regions—places where genes were in high use and coordinating cell type-specific functions.
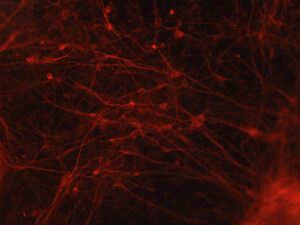
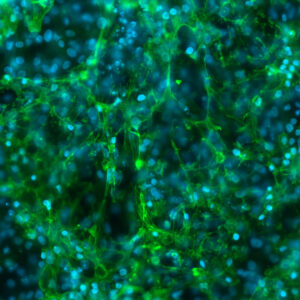
Wondering whether these dynamics may indeed influence gene expression and cell identity, the researchers moved into heart cells and neurons, deriving these from human induced pluripotent stem cells (iPSCs). They demonstrated that this dynamic organization is most important in heart cells at genes related to heart cell function and in neurons at genes related to neuronal cell function. These heart-function and neuronal-function genes are, of course, in different places in the genome—this flexibility in genome folding likely helps cell types achieve and maintain their distinct identities.
“One thing this appears to suggest is that the continuous folding and unfolding of our genome may be particularly important for helping a cell ‘remember’ who it is supposed to be by preserving expression of genes that are unique to different cell types,” says Popay.
The researchers have a few theories as to why areas of the genome related to identity seem to be the most active. Their best guess is that the constant reiteration of these loops makes identity stronger, by repeatedly creating fresh connections between genes—like the cell is constantly giving itself a pep talk, reading its affirmations in the form of proteins it needs to create to maintain its function.

Though the findings lead to new mysteries, Dixon says what they know now helps explain the symptoms associated with dysfunctional genome folding in humans.
“These genome folding machineries tightly control cell identity in every cell, so it actually makes a lot of sense that when we see mutations in them, we get these syndromic conditions like Cornelia de Lange syndrome that impact different parts of the body in different ways,” says Dixon. “And cancer is potentially exploiting that same principle, changing where in the genome these dynamics are more important to manipulate cell identity and encourage uncontrolled growth.”
With this new confirmation that the genome’s dynamic 3D structure significantly impacts gene expression, scientists can now connect the dots between genome structure and disease and begin imagining new therapies for cancers and developmental disorders. Foundational research means widespread impact—especially when it comes to the building blocks for life.
Other authors include Ami Pant, Femke Munting, Morgan Black, and Nicholas Haghani of Salk and Melodi Tastemel of UC San Diego.
The work was supported by the National Institutes of Health (U01-CA260700, S10-OD023689, S10-OD034268, P30-CA014195, P30-AG068635, P01-AG073084-04, P30-AG062429), Salk Excellerators Fellowship, Rita Allen Foundation, Pew Charitable Trusts, Howard and Maryam Newman Family Foundation, Helmsley Charitable Trust, Chapman Foundation, Waitt Foundation, American Heart Association Allen Initiative, and California Institute for Regenerative Medicine.
DOI: 10.1038/s41588-026-02516-y
JOURNAL
Nature Genetics
AUTHORS
Tessa M. Popay, Ami Pant, Femke Munting, Melodi Tastemel, Morgan E. Black, Nicholas Haghani, Jesse R. Dixon
Office of Communications
Tel: (858) 453-4100
press@salk.edu
The Salk Institute is an independent, nonprofit research institute founded in 1960 by Jonas Salk, developer of the first safe and effective polio vaccine. The Institute’s mission is to drive foundational, collaborative, risk-taking research that addresses society’s most pressing challenges, including cancer, Alzheimer's disease, and agricultural vulnerability. This foundational science underpins all translational efforts, generating insights that enable new medicines and innovations worldwide.